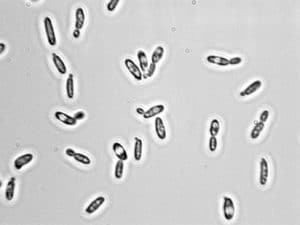
Scientists from University College of Dublin have found that two distinctly named species of yeast are in fact 99.6% identical at the base pair level, and collinear. In other words, they are the same species.
It was a bit of a shock, especially considering one of the yeast species, Pichia kudriavzevii, is commonly used in food production and classified by the US FDA as “generally recognized as safe,” while the other, Candida krusei, is known to be drug-resistant and able to cause opportunistic infections in humans.
“The existence of multiple names for this species has almost certainly impeded research into it,” the researchers write. “We suggest that P. kudriavzevii should be the only name used in future.”
Their study, published in PLOS Pathogens, highlights the importance of gathering comprehensive genetic data of organisms.
The Irish team, led by Kenneth H. Wolfe and first author Alexander P. Douglass, is the first to sequence the type strain of C. krusei. Genome sequences had been published previously for four P. kudriavzevii strains and one C. krusei clinical isolate, but they were highly fragmented, and none of them provided a chromosome-level assembly or transcriptome-based annotation.
The researchers produced high-quality reference genomes for a C. krusei type strain called CBS573 and the CBS5147 type strain for P. kudriavzevii. They then annotated the genomes with the help of RNA sequence data for CBS573, uncovering more than 5,100 protein-coding genes. They also re-sequenced 30 additional clinical and environmental isolates to explore the relationships between the strains and their genomic diversity.
Not only did the comprehensive assemblies clarify the genome content and structure, they uncovered some unexpected features of the genomes.
“One of the most unexpected features of the genome is the structure of its centromeres, which consist of a simple but large IR. The 99% DNA sequence identity of the 8–14 kb units that form the IRs means that centromere organization would have been difficult to deduce without long-read PacBio data.”
The data also allowed them to take a deeper dive into a question that has been perplexing scientists in the field concerning the sexual cycle of the yeast.
When P. kudriavzevii was first described, it was reported to be able to sporulate, forming one spore per ascus, but later studies reported that the type strain of P. kudriavzevii does not mate or sporulate.
“Our discovery that this strain is triploid provides a possible explanation for its failure to sporulate, or at least its failure to produce viable spores,” the authors write.
As for implications to health and safety, the authors say the yeast should no longer be used in food processing, as it “presents a potential hazard to the health of immunocompromised workers, and potentially also to consumers.”
They suggest that the closely related, non-pathogenic Pichia species be considered as possible alternatives for some industrial applications.