A groundbreaking new sequencer
The Onso system is an innovative benchtop short-read DNA sequencing platform with an extraordinary level of accuracy using PacBio sequencing by binding (SBB) technology. Ultra-high Q40+ data quality allows you to break through limits of detection.
15× higher accuracy than SBS sequencers
SBB chemistry delivers extraordinary levels of accuracy and sensitivity to confidently identify rare variants missed by other short-read sequencing platforms.
Less sequencing required
At Q30, the amount of coverage needed for certain applications (e.g. liquid biopsy research) is not economically feasible. The Onso system’s Q40+ performance provides high quality results at up to four fold less read depth than SBS.
Power through difficult to sequence regions of the genome
Near-perfect sequencing provides researchers accurate characterization of low complexity, highly repetitive regions for achieving more error-free genomes.
Seamless integration with existing short-read sequencing ecosystems
The Onso system is available with an accuracy-enhanced library prep solution that enables a diverse set of popular short-read applications. Plus, third-party sample and library prep solutions are supported on Onso through a simple library conversion process. The Onso FASTQ file output ensures compatibility with downstream secondary and tertiary short-read analysis tools.
Blog
Going beyond the “needle in a haystack” with Onso short-read sequencing
We’ve already shown you how the Onso system excels at needle-in-a-haystack applications, like liquid biopsy research. Now, explore how the exceptional accuracy of the Onso system extends beyond the proverbial haystack across diverse fields such as oncology, public health, and human genetics research, illustrated by three insightful case studies.
Video
SBB chemistry
Powered by innovative SBB technology, the Onso system delivers near-perfect accuracy and sensitivity to advance your research.
The system delivers 15× higher accuracy than SBS and allows characterization of homopolymers. Fewer errors mean less false positives and more biological insight.
Specifications
| Onso reagents | Read length | Reads | Output (Gb) | Run time | Quality score |
|---|---|---|---|---|---|
| 200 cycle sequencing kit |
1x200 bp |
400 — 500M (SE) |
80 — 100 | 32 hours | ≥90% Q40 |
| 300 cycle sequencing kit | 2x150 bp | 800 — 1,000M (PE) | 120 — 150 | 48 hours | ≥90% Q40 |
Download spec sheet Order kits online
What can you do with the Onso system?
Cancer research
Enable development of screening, monitoring, and therapy selection
Gene editing research
Confirmation of potential editing outcomes and biomarker discovery
Exome/panels
Validated WES using Twist Bioscience hybrid capture protocol
Single cell
10x Genomics Chromium libraries supported and validated on Onso system
Application brief
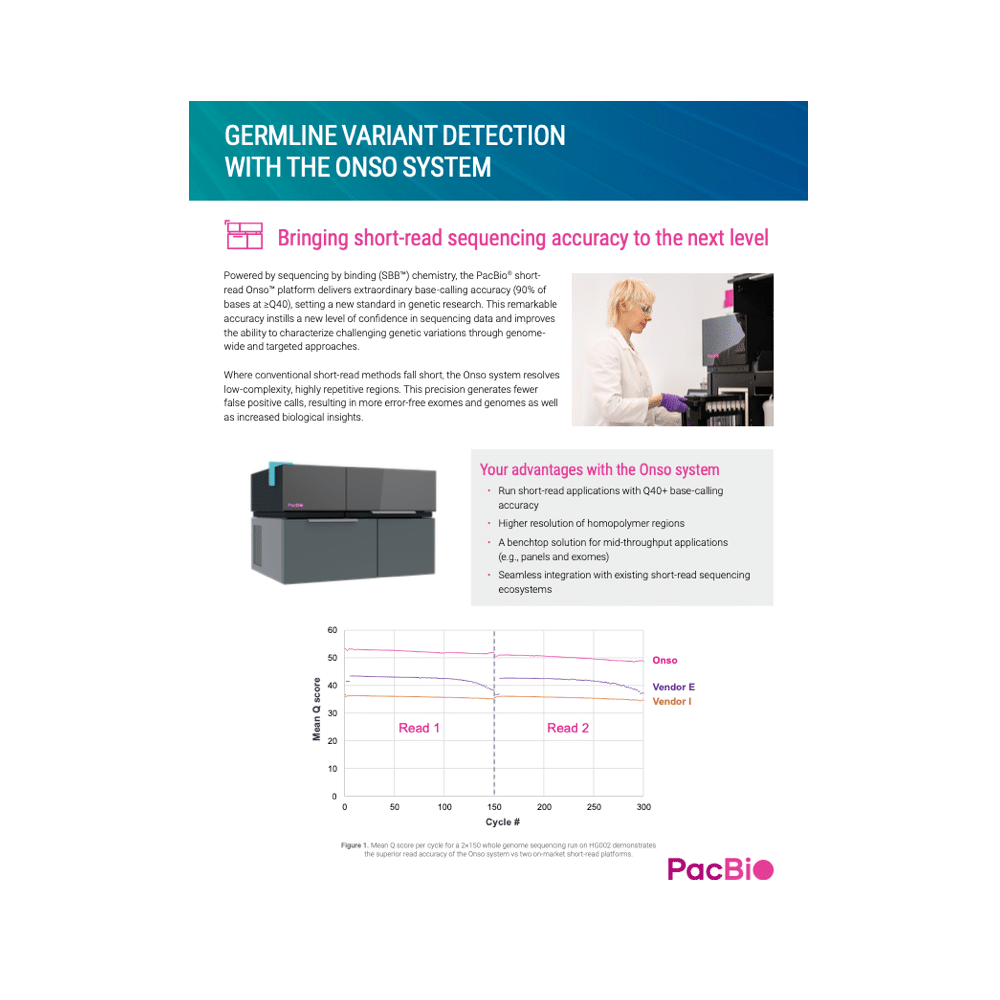
Germline variant detection with the Onso system
Powered by sequencing by binding (SBB) chemistry, the PacBio short- read OnsoTM platform delivers extraordinary base-calling accuracy (90% of bases at ≥Q40), setting a new standard in genetic research. This remarkable accuracy instills a new level of confidence in sequencing data and improves the ability to characterize challenging genetic variations through genome- wide and targeted approaches.
Industry-leading sensitivity for low-frequency allele detection
in liquid biopsy*
Variant detection performance was tested for SBB and SBS sequencing using the SeraCare complete ctDNA mutation mix across five variant allele frequencies: 0% (WT), 0.05%, 0.1%, 0.25%, and 0.5%. The Onso system achieved nearly double the sensitivity for rare variant detection with equivalent coverage (left, 6,000X SBB UMI- vs 6,000X SBS UMI-). The Onso system also demonstrated equivalent or better performance with 4-fold less sequencing (right, 6,000X SBB UMI+ vs 24,000X SBS UMI+).
PRODUCT NOTE
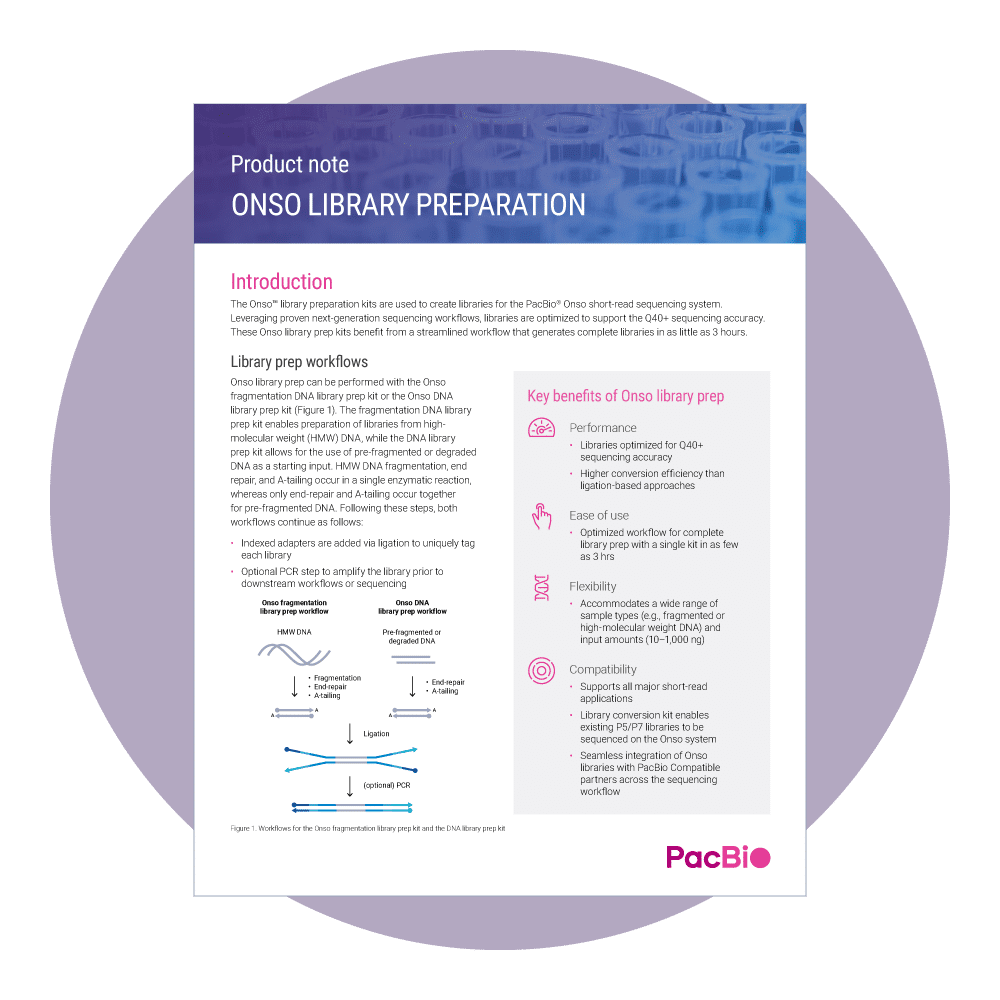
ONSO LIBRARY PREPARATION
The Onso library preparation kits are used to create libraries for the PacBio Onso short-read sequencing system. Leveraging proven next generation sequencing (NGS) workflows, libraries can be created in as little as 3 hours from a wide range of sample types (e.g., fragmented or high-molecular weight DNA) and input amounts (10–1,000 ng).
Frequently asked questions
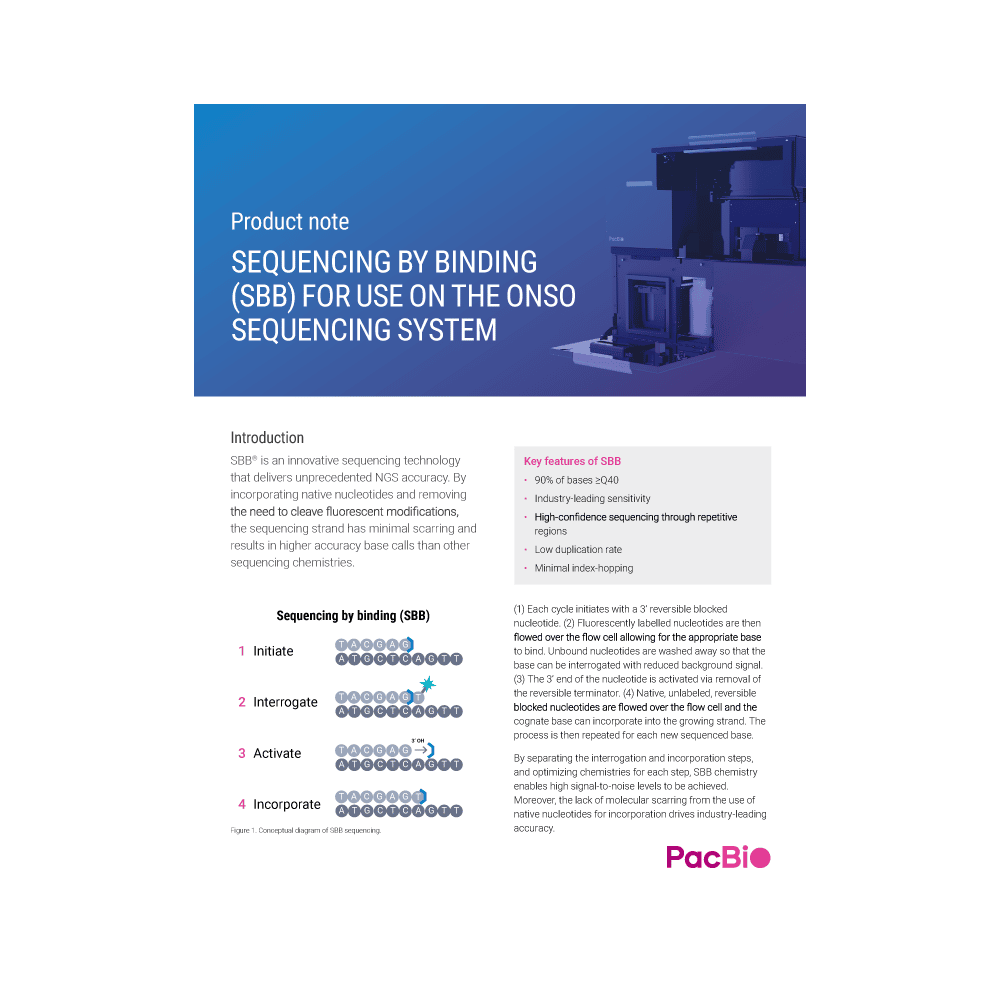
Separation of the interrogation (detection) and incorporation (base extension) steps, along with optimized chemistries tuned for these steps, allow us to achieve raw accuracies of Q40+. In addition, use of our library prep to minimize end-repair errors allows our platform to achieve upwards of Q50.
WGS: Approximately 1 sample (2×150 PE)
Exomes: Assuming 100x mean depth of coverage / 42Mb exome size running 2×100 PE can accommodate up to 21 exomes
scRNA-Seq: Assuming 5,000 cells and 25,000 read pairs/cell, a 28×90 PE run can accommodate 4 samples
The Onso system sequencing instrument includes a computer (APC) for performing on-board primary analysis basecalling, demultiplexing, and FASTQ file generation. Demultiplexing and FASTQ generation can also be performed off-instrument by running a command line software tool provided by PacBio for customer’s own compute system(s)
At this time, no secondary or tertiary solutions will be offered with the Onso system; however, we anticipate Onso FASTQ datatypes should be compatible with the existing secondary and tertiary solutions available for short-read NGS platforms
The Onso system was designed with flexibility to allow users to leverage the large number of genomic tools available for short-read NGS. Libraries prepared from third party kits are easily adapted for use on the Onso system with the Onso library conversion kit. This enables applications including single-cell analysis, exome sequencing and general targeted panel sequencing using hybrid capture technology.
PacBio Compatible products
Power your Onso with PacBio Compatible partners
The PacBio Compatible program provides seamless integration of industry-leading tools into the short-read sequencing by binding (SBB) workflow on the Onso platform.
Higher accuracy
Unique SBB technology delivers 15× higher accuracy for sensitive and specific characterization of low complexity regions with 90% Q40+. Fewer errors mean less false positives and more biological insight.
Improved sensitivity
Using fewer reads, the Onso system lowers the limit of detection, which drives the sensitivity of applications like liquid biopsy research. This enables the identification of variants missed by other short-read sequencing platforms.
Streamlined workflow
Simple workflows easily integrate into existing laboratory processes, reducing time and labor.
Efficiency
Q40+ delivers high quality results at lower read depths enabling scalable sequencing while reducing time and cost. Reduced need for unique molecular identifiers (UMIs) lowers workflow complexity and cost.
Product note
SEQUENCING BY BINDING (SBB) FOR USE ON THE ONSO SEQUENCING SYSTEM
SBB is an innovative sequencing technology that delivers amazing NGS accuracy. By incorporating native nucleotides and removing the need to cleave fluorescent modifications, the sequencing strand has minimal scarring and results in higher accuracy base calls.
This is your moment
Our highly differentiated portfolio of sequencing systems provide the tools you need to accelerate your research. From the fullest picture to the finest detail, your moment of discovery is waiting.