
Genetic knowledge is powerful when it comes to breeding. The ability to trace desirable traits to the gene level can help create plants and animals that are adapted to existing and emerging challenges, such as temperature tolerance, productivity, or disease resistance.
By crossing two breeds of cattle, Angus (Bos taurus taurus) and Brahman (Bos taurus indicus), from opposite ends of the species spectrum, breeders can benefit from the Angus’s high productivity in cool environments and the Brahman’s tolerance for harsh, hot climates and the diseases and parasites found there.
Genetically and phenotypically, the two subspecies are very different. And, their offspring are as well, as John Williams of the University of Adelaide explained in his recent talk at PAG. There are even differences depending on how the breeds are crossed (i.e. Angus bull and Brahman cow, or Brahman bull and Angus cow). Fetal weight at mid-gestation, for instance, varies markedly among purebreds and crosses, and between crosses.
Interested in exploring these differences, Williams and colleagues embarked on two approaches to assembling this heterozygous genome.
The first was a one-technology methodology involving PacBio long-read sequencing, assembled with FALCON-Unzip and scaffolded with Hi-C data, to examine the genome of an F1 cross-breed (Angus x Brahman). The Iso-Seq method was then used to explore the cattle’s transcriptome. It enabled the team to examine entire transcripts, as well as isolate 30,000 isoforms from 12,000 genes.
Although they are still sifting through the data, Williams said the team is “starting to be able to differentiate between Angus and Brahman specific transcripts.”
“Initial results show that Iso-Seq data can be haplotyped and is highly concordant with genome phasing results, revealing possible allelic-specific isoform expression,” he added.
Mapped back to the assembly, the Iso-Seq data also confirmed that the F1 cattle reference genome is of good quality.
Among the genes they explored, 10 were heavily differentially expressed between male and female. The team wanted to drill down deeper, to determine which parent of origin the differences come from, and to create better assemblies of sex specific genes.
So, a sub group led by Adam Phillippy, Sergey Koren, and Arang Rhie of the National Human Genome Research Institute (NHGRI) in Bethesda, Maryland, created a new process that took advantage of access to the cattle’s parents.
Trio Binning: Two Genomes From One Individual
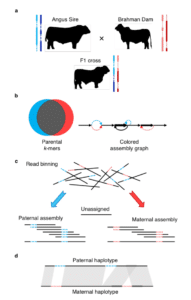 The “trio binning” process, also presented at PAG, enabled them to generate two high-quality (maternal and paternal) genomes from the single F1 cross-breed. It uses short reads from two parental genomes to partition SMRT Sequencing long reads from an offspring into haplotype-specific sets prior to assembly. Each haplotype is then assembled independently using a new module of the Canu assembler the NHGRI team created — TrioCanu — resulting in a complete diploid reconstruction.
The “trio binning” process, also presented at PAG, enabled them to generate two high-quality (maternal and paternal) genomes from the single F1 cross-breed. It uses short reads from two parental genomes to partition SMRT Sequencing long reads from an offspring into haplotype-specific sets prior to assembly. Each haplotype is then assembled independently using a new module of the Canu assembler the NHGRI team created — TrioCanu — resulting in a complete diploid reconstruction.
As described in this preprint, the method requires moderate coverage of short sequencing reads (e.g. 30-fold Illumina) from two parental genomes to identify short, k-length subsequences (k-mers) that are specific to each parent. These k-mers are presumed to be specific to the corresponding haplotypes of the offspring. Next, long reads are collected from an offspring of the parents to sufficiently cover both haplotypes (e.g. 80-fold PacBio, 40-fold per haplotype). Long reads from the offspring are then binned into paternal and maternal groups based on the presence of the haplotype-specific k-mers and assembled separately.
In the case of the cattle, the Angus and Brahman haplotypes aligned to one another with 99.35% identity and contained 25,245 haplotype-specific structural variants and 124 inversion breakpoints.
Phillippy et al. note that trios have long been used in genomics to infer inheritance, including for the HapMap and the 1000 Genomes projects, as well as by trio-sga to simplify heterozygous diploid genome assembly. But reliance on short-read sequencing limited the haplotype-specific contigs (haplotigs) to an average size of a few kilobases.
“In contrast, our long-read method enables the assembly of multi-megabase haplotigs and complete parental haplotypes,” the authors write.
Long-read trio binning is also advantageous because it requires fewer resources than inbreeding, simplifies assembly graphs, and can accurately reconstruct structurally heterozygous alleles that can be important factors in adaptation and immunity, Phillipy states.
“Accurate representation of haplotypes is essential for studies of intraspecific variation, chromosome evolution, and allele-specific expression,” the authors add.
Its applications could also spread into human and other areas of agricultural genomics, including polyploid plant genomes.
“Reference genome projects have historically selected inbred individuals to minimize heterozygosity and simplify assembly,” the authors write. “We challenge this dogma and present a new approach designed specifically for heterozygous genomes.”