 Single Molecule, Real-Time (SMRT) Sequencing continues to get smarter and more powerful, with the recent launch of the Sequel II system increasing capabilities and efficiencies of the long-read DNA and RNA PacBio sequencing technology even further. In a special issue devoted entirely to the technology in the MDPI open access journal Genes, guest editors Adam Ameur of Uppsala University and Matthew S. Hestand of the Cincinnati Children’s Hospital Medical Center present eight articles highlighting research conducted using SMRT Sequencing.
Single Molecule, Real-Time (SMRT) Sequencing continues to get smarter and more powerful, with the recent launch of the Sequel II system increasing capabilities and efficiencies of the long-read DNA and RNA PacBio sequencing technology even further. In a special issue devoted entirely to the technology in the MDPI open access journal Genes, guest editors Adam Ameur of Uppsala University and Matthew S. Hestand of the Cincinnati Children’s Hospital Medical Center present eight articles highlighting research conducted using SMRT Sequencing.
As this special issue demonstrates, the benefits of SMRT Sequencing to many different areas of research are becoming evident, not only in basic science, but also in more applied areas such as agricultural, environmental, and medical research. Examples from each of these areas are included in this issue.
Maximizing Minimum Sample Sizes
A new mosquito genome assembly generated via a collaboration between PacBio and the Wellcome Sanger Institute highlighted the capabilities of one of the latest SMRT Sequencing advancements: a new low DNA input protocol. The Sanger scientists used the new protocol to create a high-quality de novo genome assembly from a single Anopheles coluzzii mosquito. A modified library construction protocol without DNA shearing and size selection was used to generate a SMRTbell library from just 100 ng of starting genomic DNA.
Protecting the Fungus Among Us

Scientists in China and Michigan used SMRT Sequencing to elucidate the medicinal properties of Gloeostereum incarnatum, a precious edible mushroom that is widely grown in Asia. They assembled a high-quality genome of the fungus — the first complete genome to be sequenced in the family Cyphellaceae — and identified gene clusters associated with terpenoid and polysaccharide biosynthesis.
Another team from Jilin Agricultural University and Shenyang Agricultural University in China also investigated edible mushrooms — and a mycoparasite that threatens them. They assembled a high-quality genome of Cladobotryum protrusum, which causes cobweb disease on cultivated mushrooms. They found that the C. protrusum genome, the first complete genome to be sequenced in the genus Cladobotryum, encodes a large and diverse set of genes involved in pathogen–host interactions, mycotoxins, and pigments, and harbors arrays of genes with the potential to produce bioactive secondary metabolites and stress response-related proteins that are significant for adaptation to hostile environments.
Improving Protein Production… via Insects

A new genome assembly of the cabbage looper moth, Trichoplusia ni, may have implications for large-scale genome engineering. As reported by scientists from the National Cancer Institute’s Frederick National Laboratory for Cancer Research, insect cell protein production has emerged as a viable alternative to bacterial and mammalian cells for the production of therapeutically relevant proteins, with several approved vaccines generated in baculovirus-infected insect cells. However, improved protein production using these lepidopteran hosts has been hindered by limited genomic data. By performing de novo genome assembly of the Trichoplusia ni-derived cell line Tni-FNL, the team hopes the reference will bolster future large-scale genome engineering work in recombinant protein production hosts.
Detecting Distinction in Bone Marrow Subpopulations
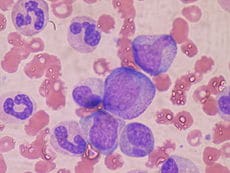 In a study led by Anne Deslattes Mays and Anton Wellstein from the Lombardi Comprehensive Cancer Center at Georgetown University, the transcriptomes of freshly harvested human bone marrow progenitor (lineage-negative) and differentiated (lineage-positive) cells were analyzed with SMRT full-length RNA sequencing. This Iso-Seq analysis revealed a ~5-fold higher number of transcript isoforms than previously detected and showed a distinct composition of individual transcript isoforms characteristic for bone marrow subpopulations. Check out an additional Q&A with Mays here.
In a study led by Anne Deslattes Mays and Anton Wellstein from the Lombardi Comprehensive Cancer Center at Georgetown University, the transcriptomes of freshly harvested human bone marrow progenitor (lineage-negative) and differentiated (lineage-positive) cells were analyzed with SMRT full-length RNA sequencing. This Iso-Seq analysis revealed a ~5-fold higher number of transcript isoforms than previously detected and showed a distinct composition of individual transcript isoforms characteristic for bone marrow subpopulations. Check out an additional Q&A with Mays here.
Expanding Genetic Diversity in Human Dataset
Swedish scientists used SMRT Sequencing to expand the diversity of the human genome dataset. They performed de novo assembly of two Swedish genomes that revealed over 10 Mb of sequences absent from the human GRCh38 reference in each individual, and around 6 Mb of novel sequences (NS) shared with a Chinese personal genome. “Our results highlight that the GRCh38 reference is not yet complete and demonstrate that personal genome assemblies from local populations can improve the analysis of short-read whole-genome sequencing data,” the authors wrote.
Solving Methylation Calling Challenges
Lastly, a team of bioinformaticians from Iowa, Ohio, and Tokyo presented a statistical solution for observing personal diploid methylomes and transcriptomes. As they report, CpG methylome pairs of homologous chromosomes that are distinguishable with respect to phased heterozygous variants (PHVs) is challenging due to scarcity of PHVs in personal genomes. While SMRT Sequencing is a promising avenue to addressing this challenge as it outputs long reads with CpG methylation information, phasing the CpG sites can still come with errors. Their paper proposes a model that reduces the error rate to 1%, thereby calling CpG hypomethylation in each haplotype with >90% precision and sensitivity.