It’s a moment three decades in the making: the first complete human genome assembly is here!
Reading this you will no doubt feel some sense of déjà vu. After all, the human genome reference was pronounced “done” in 2000, 2001, and again in 2003. But any scientist who has used the reference since then knows that there has never been a single fully sequenced human genome. Until now.
HiFi sequencing enables the first complete sequence of a human genome
The Telomere-to-Telomere (T2T) Consortium, a large team of scientists from the National Human Genome Research Institute and dozens of other institutions, released a new preprint titled “The complete sequence of a human genome.” Lead authors Sergey Nurk, Sergey Koren, Arang Rhie, and Mikko Rautiainen, along with corresponding authors Evan Eichler, Karen Miga, and Adam Phillippy as well as many collaborators have now vanquished gaps and errors to deliver what they call “the first truly complete human reference genome.”
This tremendous effort incorporated several cutting-edge technologies, including HiFi sequencing from PacBio, to produce a gap-free, complete haploid human genome assembly based on a complete hydatidiform mole (CHM13). The goal was to create a novel resource with comprehensive, reliable genome data that avoids the gaps and errors that still mark the latest GRCh38 reference assembly. “The resulting T2T-CHM13 reference assembly removes a 20-year-old barrier that has hidden 8% of the genome from sequence-based analysis, including all centromeric regions and the entire short arms of five human chromosomes,” Nurk et al. report.
New to HiFi sequencing? – Learn about this new paradigm in sequencing technology.
This new reference “includes gapless assemblies for all 22 autosomes plus chromosome X, corrects numerous errors, and introduces nearly 200 million bp of novel sequence containing 2,226 paralogous gene copies, 115 of which are predicted to be protein coding,” the authors add. This represents “the largest improvement to the human reference genome since its initial release.”
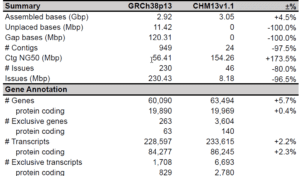
HiFi sequencing was pivotal to this achievement. The scientists note that HiFi sequencing features “20 kbp read lengths and a median accuracy of 99.9%, which has resulted in unprecedented assembly accuracy with relatively minor adjustments to standard assembly approaches. …HiFi sequencing excels at differentiating subtly diverged repeat copies or haplotypes.”
HiFi sequencing removes technological barriers
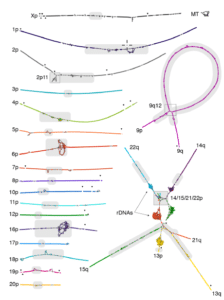
The team had initially started with a strategy of using noisy ultralong nanopore-based reads to build an assembly backbone, which was then polished with other platforms. But they subsequently switched to accurate and long HiFi reads. “We shifted to a new strategy that leverages the combined accuracy and length of HiFi reads to enable assembly of highly repetitive centromeric satellite arrays and closely related segmental duplications,” they report. The assembly is based on a string graph built from HiFi reads and has an average consensus accuracy between Q67 and Q73, “far exceed[ing] the original Q40 definition of ‘finished’ sequence,” the authors add.
The new assembly, to which a Y chromosome sequence will be added in the near future, should be used in place of the GRCh38 reference for “all studies requiring a linear reference sequence,” the scientists suggest, noting that it is “more complete, representative, and accurate” than its predecessor and “substantially increases the number of known genes and repeats in the human genome.”
The team also notes that reanalysis of short-read public data sets such as the 1000 Genomes Project using the new reference already shows improvement compared to the GRCh38 reference, and that new phenotypic associations should be expected given the more complete reference genome.
HiFi sequencing powers the next phase of genomic discovery
“The complete, telomere-to-telomere assembly of a human genome marks a new era of genomics where no region of the genome is beyond reach,” the authors write.
“Highly accurate, long-read sequencing, combined with tailored algorithms, promises the de novo assembly of individual haplotypes and sequence-level resolution of complex structural variation. This will require the routine and complete de novo assembly of diploid human genomes, as planned by the Human Pangenome Reference Consortium.”
Ultimately, they anticipate that highly accurate long-read sequencing will lead to a “collection of high-quality, complete reference haplotypes [that] will transition the field away from a single linear reference and towards a reference pangenome that captures the full diversity of human genetic variation,” the team reports. “Ideally, every genome could be assembled at the quality achieved here, since the small variants recovered by short-read resequencing approaches represent only a fraction of total genomic variation.”
How to get started with HiFi sequencing for any genome
- Learn more about our whole genome sequencing application.
- Have your questions about HiFi sequencing answered by a PacBio scientist.
- 2021 HiFi for Accuracy SMRT Grant program — Apply between June 7–25 for your chance to win free HiFi sequencing.