 According to many, PacBio is the new “gold standard” in microbial sequencing. Chief Scientific Officer Jonas Korlach notes that its ability to simultaneously provide long sequencing reads (genome contiguity), high consensus accuracy (genome accuracy), minimal sequence bias (genome completeness), and methylation detection (bacterial epigenome) has made it the technology of choice for users who need to reliably produce high quality genomes.
According to many, PacBio is the new “gold standard” in microbial sequencing. Chief Scientific Officer Jonas Korlach notes that its ability to simultaneously provide long sequencing reads (genome contiguity), high consensus accuracy (genome accuracy), minimal sequence bias (genome completeness), and methylation detection (bacterial epigenome) has made it the technology of choice for users who need to reliably produce high quality genomes.
In a presentation for the virtual Microbiology & Immunology conference, Korlach highlighted PacBio’s strengths in the field, including multiplexed microbial sequencing on the Sequel System and full-length bacterial RNA sequencing.
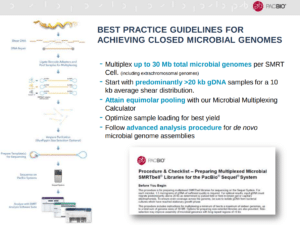 Microbial de novo genome assembly
Microbial de novo genome assembly
Multiple bacterial genomes can now be sequenced in one SMRT Cell. Not only are bacterial chromosomes revealed, but plasmids are also assembled. This is significant because these mobile genetic elements often carry the genes that drive virulence, drug resistance, and other traits that are important to understand microbial biology, Korlach said.
Getting the full picture of bacterial plasmids allows scientists to track the transmission and follow hospital-associated infection outbreaks, for instance. It also gives insights into the evolution of bacterial strains, both through single nucleotide changes and larger structural rearrangements.
“It is now possible for the first time to really understand the evolution and generation of some of these very dangerous superbugs that are resistant to all known antibiotics,” Korlach said, citing a study of 16 Klebsiella pneumoniae isolates collected in German hospitals.
Korlach also referenced a study that solved a decades-old mystery about Spiroplasma poulsonii, a type of symbiotic bacteria that manipulate host reproduction to spread in a population by selectively killing off the sons of infected female hosts during development. A team of Swiss researchers identified the toxin responsible, located on a plasmid, using SMRT Sequencing.
“This thorough understanding of the functional ramifications was really only provided by PacBio sequencing, and forms the foundation now for thinking about controlling insect populations.”
Other advantages of PacBio sequencing? Every bacteria has multiple copies of the 16S housekeeping gene, and PacBio is the only technology able to produce multiple distinct 16S sequences per bacterial genome, Korlach said. PacBio sequencing can also overcome long-standing challenges in assembling yeast, fungi and other eukaryotic genomes connected to infectious disease.
Malaria causing agent plasmodium falciparum, for instance, was nearly impossible to sequence in traditional platforms because it is highly repetitive and AT-rich. But PacBio long-read sequencing enabled complete telomere-to-telomere de novo assembly of the genome. Another recent publication highlighted the utility of PacBio sequencing for closing yeast genomes, revealing that differently named food processing and pathogenic strains of yeast are in fact the same species.
Characterizing the bacterial epigenome and transcriptome
One of the three pillars of understanding bacterial biology is the epigenome, which PacBio is uniquely suited to characterize without any additional sequencing beyond what is needed for genome assembly. Characterizing methylomes and methylation status can shed light on how some bacteria can switch pathogenicity between not-so-dangerous to fatal, or the changes that occur when free-living bacteria associate with a host and become symbiotic instead.
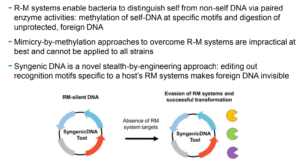
Researchers at The Forsyth Institute have created a method for transforming formerly resistant bacterial targets by leveraging the epigenetic fingerprinting information revealed by SMRT Sequencing.
Their SyngenicDNA stealth-based evasion of restriction-modification barriers during bacterial genetic engineering could unlock myriad applications in basic research, industrial biology, synthetic biology, and translational science, Korlach said.
Finally, Korlach referenced a collaboration with New England BioLabs to adapt the Iso-Seq full-length RNA Sequencing protocol to bacterial transcripts, providing the first detailed look at the dynamic structure and regulation of operon transcription.
“It was an unprecedented view of the complexity of the bacterial transcriptome and bacterial gene expression that was previously hidden.” Korlach said.
Check yourself
 Korlach also delivered a warning: Relying on existing NCBI database entries to establish what you have in your lab may be risky.
Korlach also delivered a warning: Relying on existing NCBI database entries to establish what you have in your lab may be risky.
“A number of the so-called complete genomes in the NCBI database that were done a few years ago contain, in some cases, quite dramatic errors,” he said.
A survey of 20 “reference strains” contained in the GenBank found that 30% had significant structural alterations when re-sequenced and assembled from scratch using PacBio technology, either due to previous sequencing errors or bacterial changes while in the repository.
“If you really want to be sure about what you have in your tube or on your plate, I suggest doing the genome from scratch,” Korlach said.
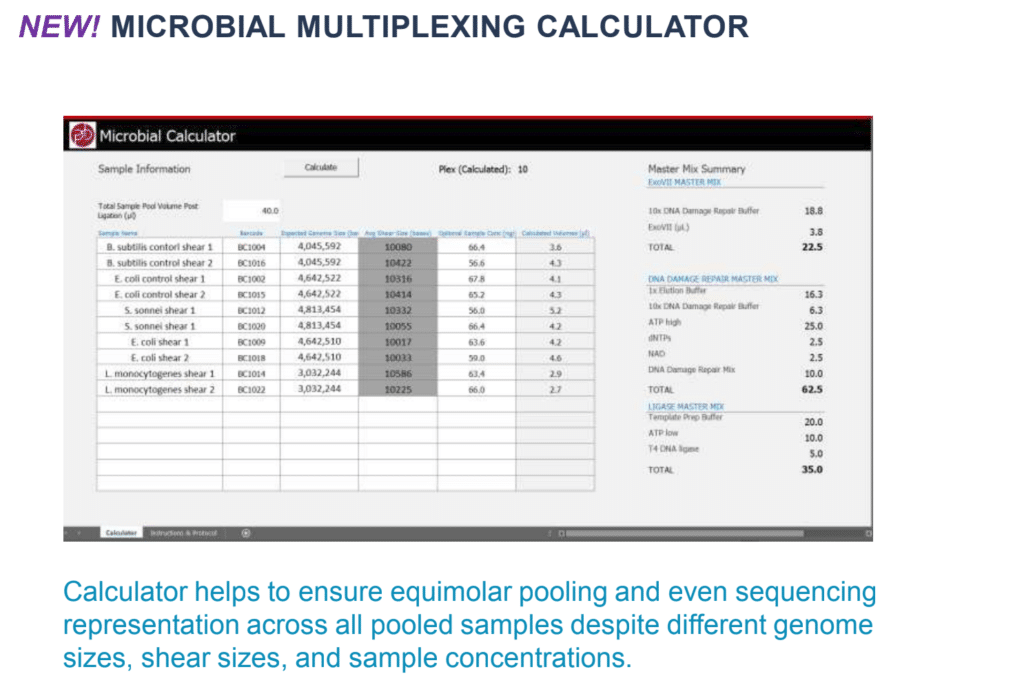
Luckily, this has become easier and more accessible with microbial multiplexing. Korlach outlined some of the nuts and bolts of the new microbial multiplexing kit, released in June, that allows researchers to pool numerous bacteria (of around 30 Mb) in one sample and sequence it in one reaction. He also pointed people to a microbial multiplexing calculator to help ensure equimolar pooling and even sequencing representation across all pooled samples despite different genome sizes, shear sizes, and sample concentrations.
Watch the complete presentation or check out additional examples of how scientists are advancing our understanding of microbial genomes, epigenomes, and communities with SMRT Sequencing.
And those interested in attending and presenting at ASM Microbe in San Francisco, June 20-24, should note that the deadline for abstract submissions has been extended to Jan. 25, 2019.