HIFI SEQUENCING
DELIVERING HIGHLY ACCURATE LONG READS TO DRIVE DISCOVERY IN LIFE SCIENCE
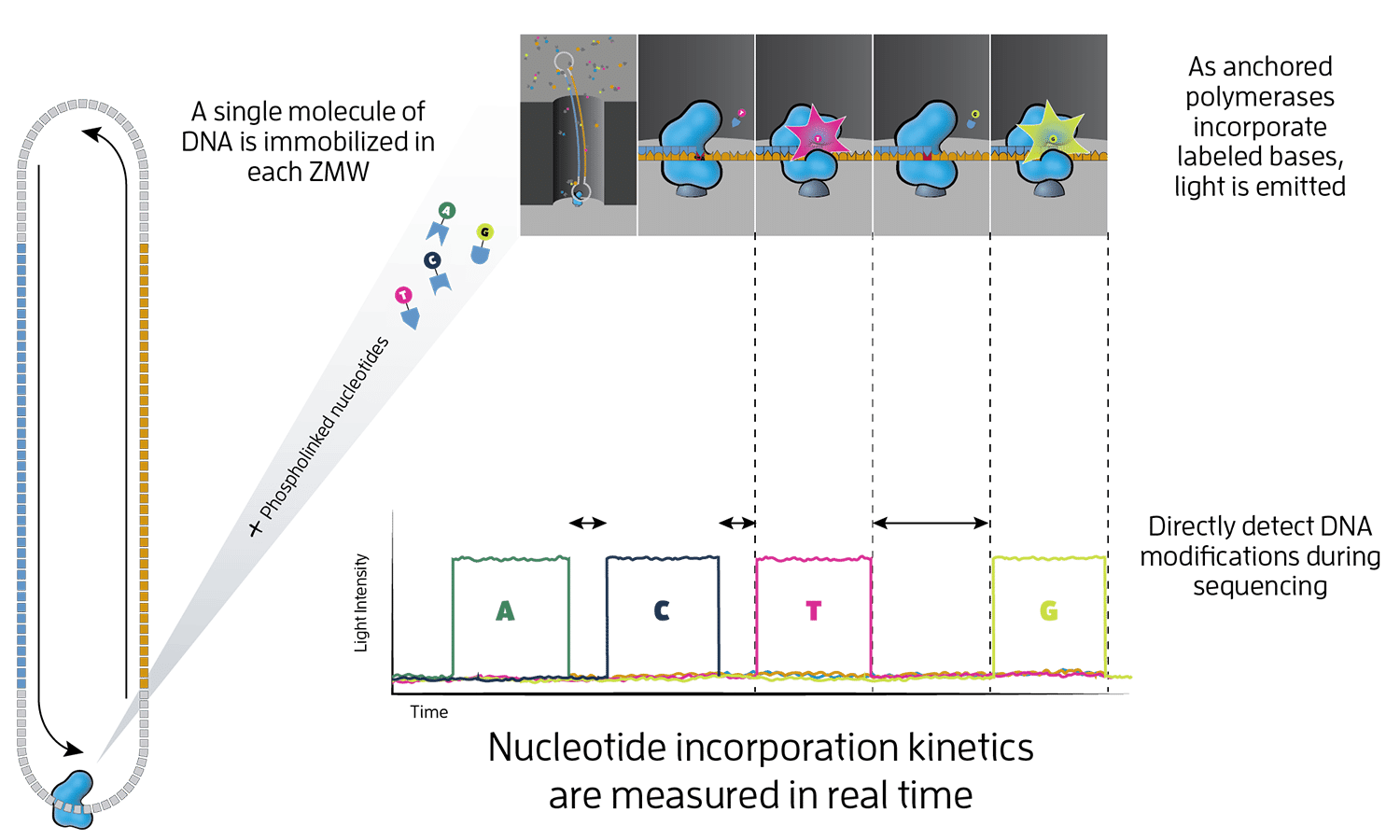
What is HiFi sequencing?
HiFi sequencing is the core technology powering our long-read sequencing platforms. This innovative approach was the first of its kind and is now a proven technology used in all fields of life science.
Original publication: Eid, J., et al. (2009) Real-time DNA sequencing from single polymerase molecules. Science, 323(5910), 133–138.
What are the advantages of HiFi sequencing?
Long reads
With reads tens of kilobases in length you can readily assemble complete genomes and sequence full-length transcripts. HiFi sequencing provides exceptional read lengths without compromising throughput or accuracy.
Data from a 15 kb size-selected human library using the SMRTbell express template prep kit 2.0 on a Sequel IIe system (2.0 chemistry, Sequel IIe system software v10, 30-hour movie). Read lengths, reads/data per SMRT Cell, and other sequencing performance results vary based on sample quality/type and insert size.
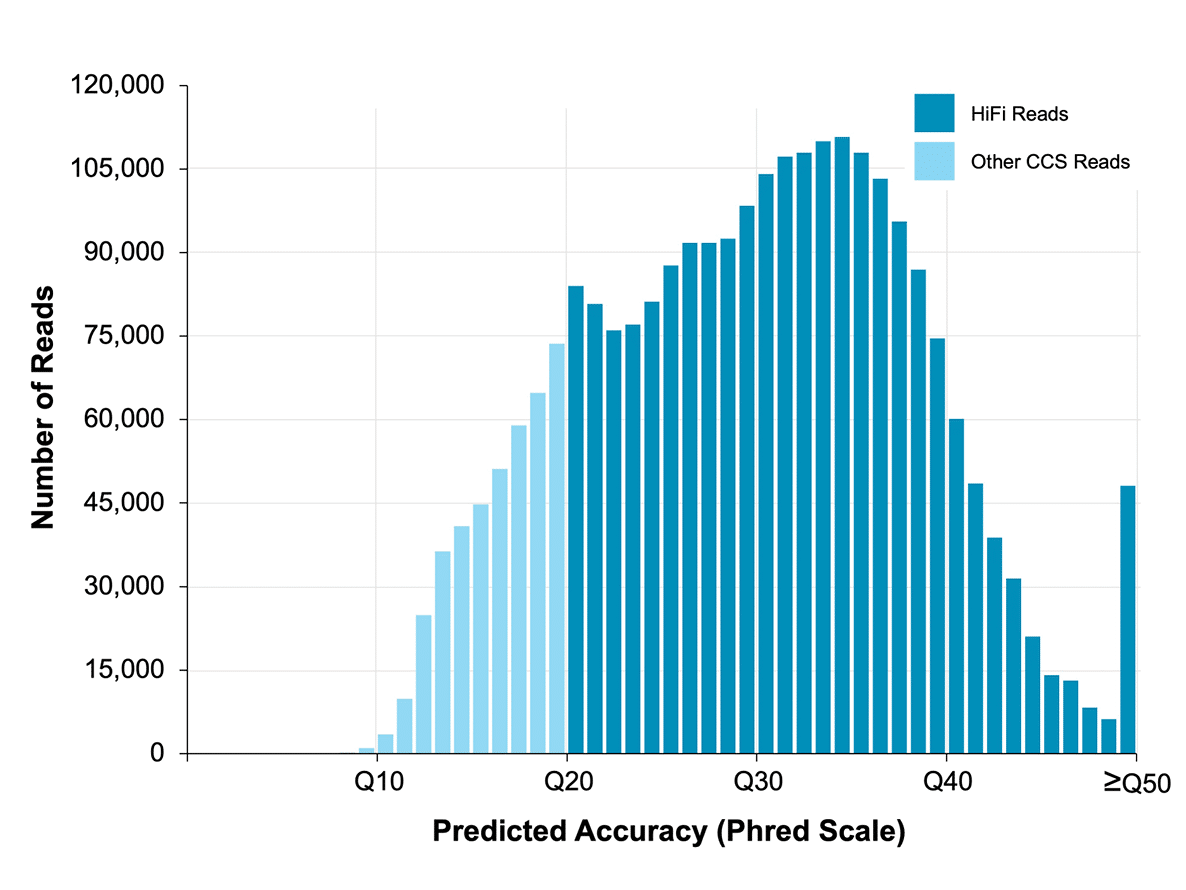
High accuracy
Sequencing free of systematic error achieves >99.999% consensus accuracy. Explore the benefits of highly accurate long-read sequencing. With exceptional accuracy, low sequencing-context bias, and accurate mapping of reads, HiFi sequencing provides the information needed to confidently call and detect all variants.
Data from a 15 kb size-selected human library using the SMRTbell express template prep kit 2.0 on a Sequel IIe system (2.0 chemistry, Sequel IIe system software v10, 30-hour movie). Read lengths, reads/data per SMRT Cell, and other sequencing performance results vary based on sample quality/type and insert size.
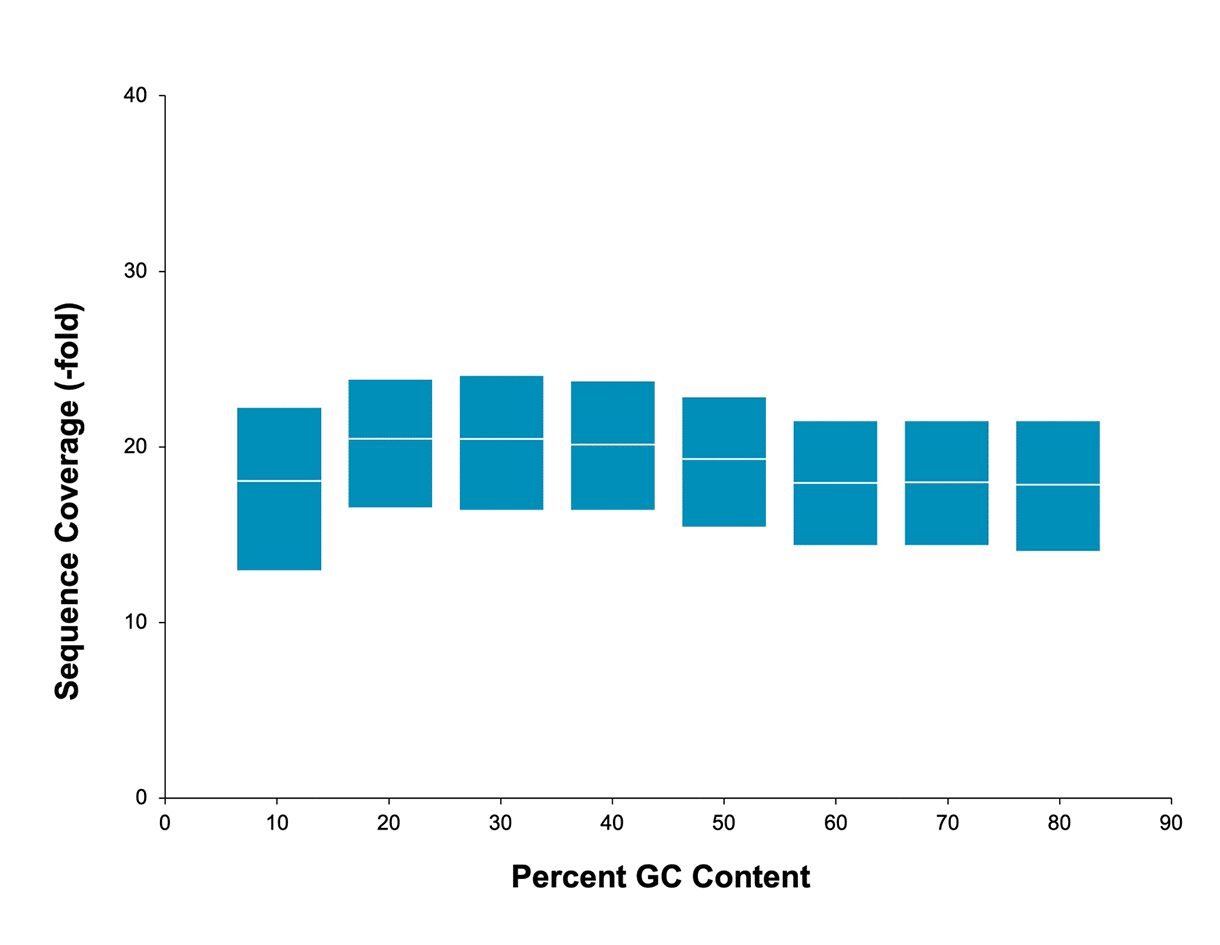
Uniform coverage
No bias based on GC content means you can sequence through regions inaccessible to other technologies. Readily sequence through AT-rich or GC-rich regions, highly repetitive sequences, long homopolymers, and palindromic sequences with HiFi sequencing.
Mean coverage per GC window across a human sample. Data generated with a 20 kb HiFi library on a Sequel II system (2.0 chemistry and Sequel II system). Read lengths, reads/data per SMRT Cell, and other sequencing performance results vary based on sample quality/type and insert size.
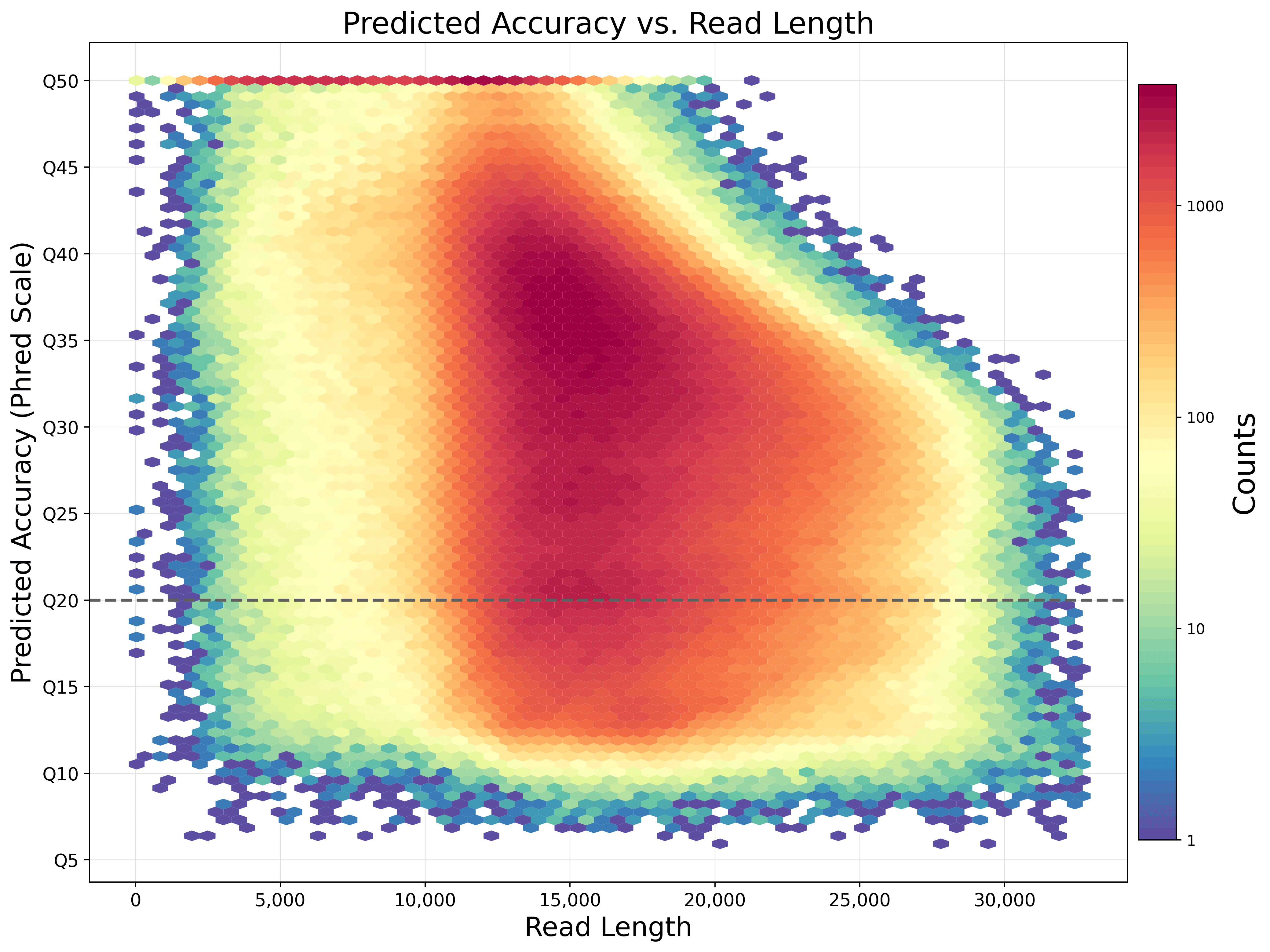
Single-molecule resolution
Capturing sequence data from native DNA or RNA molecules enables highly accurate long reads with >99.9% single-molecule accuracy. Readily achieve high-quality reads to confidently resolve variants of all types.
Data from a 15 kb size-selected human library using the SMRTbell express template prep kit 2.0 on a Sequel IIe system (2.0 chemistry, Sequel IIe system software v10, 30-hour movie). Read lengths, reads/data per SMRT Cell, and other sequencing performance results vary based on sample quality/type and insert size.
Epigenetics
With no PCR amplification step, base modifications are directly detected during sequencing. Measurement of variation in polymerase kinetics of DNA base incorporation eliminates the need for chemical modification to detect base modifications. This allows you to capture sequence and epigenetic information in a single experiment.