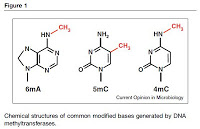
A new review paper from Brigid Davis, Michael Chao, and Matthew Waldor at Harvard Medical School considers a number of recent studies and findings that have used single molecule, real-time (SMRT®) sequencing to generate epigenomic information. “Entering the era of bacterial epigenomics with single molecule real time DNA sequencing” was recently published in Current Opinion in Microbiology.
In the review, the authors note the importance of fully understanding and analyzing genome-wide methylation data, but say that technologies to date have not made it feasible to generate this information. “The advent of new sequencing platforms in the last decade has allowed the pace of whole genome sequencing to increase exponentially,” they write. “Meanwhile, large scale analyses of DNA modification have lagged far behind, resulting in a widening gap between the extents of genomic and epigenomic knowledge.”
With the availability of SMRT Sequencing, the only sequencing platform that simultaneously gathers base modification data while sequencing the DNA, that gap may now begin to shrink considerably. The authors note, “This technological advance sets the stage for comprehensive, mechanistic assessment of the effects of bacterial DNA methyltransferases (MTases) — which are ubiquitous, extremely diverse, and largely uncharacterized — on gene expression, chromosome structure, chromosome replication, and other fundamental biological processes.”
Waldor and his colleagues review a number of recent studies that have utilized SMRT Sequencing for characterizing bacteria, noting that in many cases the findings were unexpected and revealed a significant amount of new information that could not have been predicted from DNA sequence alone. They point to the well-studied Dam methyltransferase, reporting that this epigenetic mark has been linked to the start of DNA repair and chromosome replication as well as to the modulation of gene expression and virulence in pathogens.
In the future, the authors note, merging interpretations from methylome data with other genome-wide information will become increasingly critical to analyze bacteria in context for a comprehensive view of biological processes. “Epigenomic profiling is likely to be particularly powerful when coupled with additional investigative approaches such as transcriptome analyses,” they write.
The authors state, “we are now poised to begin an exciting era of characterizing the roles of DNA modifications throughout the bacterial kingdom.”
And while we’re on the subject, if microbial genomics is up your alley, don’t miss our workshop on epigenomes of microbes at the ASM annual meeting next month (Saturday, May 18, 1:00 pm). It’s workshop 24, “Studying Whole-Genome Microbial Epigenetics” and you can register for it as you sign up for the conference.